SRA: Sequence Read Archive
The Sequence Read Archive (SRA) makes biological sequence data available to the research community to enhance reproducibility and allow for new discoveries by comparing data sets. The SRA stores raw sequencing data and alignment information from high-throughput sequencing platforms, […]
However, it is rather difficult to even find the download links… and even then…
This web site “SRA-explorer” makes things easier and provides pre-edited commands.
EVEN THOUGH, missing SRA folders are still sometimes included…
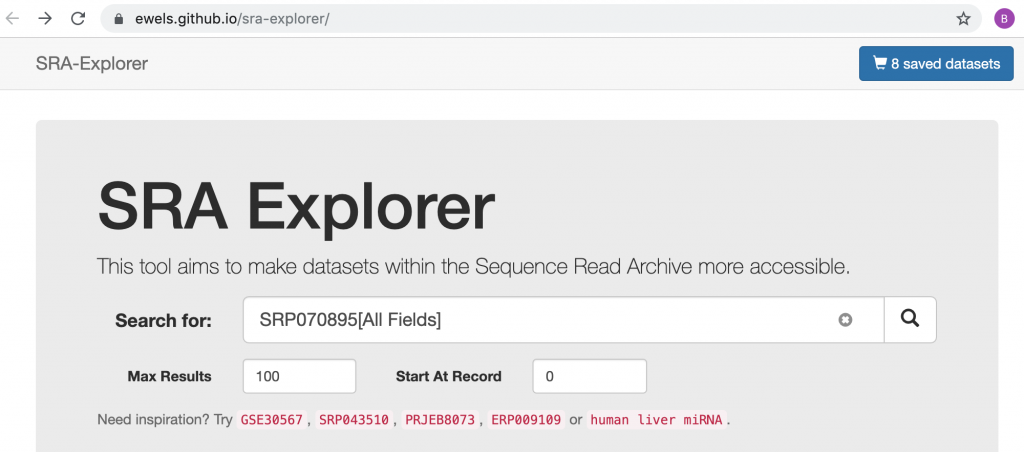
Here’s how I ended-up there….
- It all started with trying a “Dockerised” RNA-Seq pipeline: An open RNA-Seq data analysis pipeline tutorial with an example of reprocessing data from a recent Zika virus study. (Wang Z, Ma’ayan A. PubMed: 27583132)
- Docker Hub entry maayanlab/zika
- They propose a link to a “Project”: ftp://ftp-trace.ncbi.nlm.nih.gov/sra/sra-instant/reads/ByStudy/sra/SRP/SRP070/SRP070895/
- The script just would need this Project ID to download all 8 SRA files.
- BUT this is no longer on the SRA ftp site…
- After lots of searches I found the SRA-explorer web site that provided exact links for each experiment: For example:
ftp://ftp-trace.ncbi.nlm.nih.gov/sra/sra-instant/reads/ByRun/sra/SRR/SRR319/SRR3191542/SRR3191542.sra - OK… BUT the folder
SRR3191542no longer exists on the ftp site… neither areSRR3191543,SRR3191544, orSRR3191545 - Thanks to SRA-explorer it is possible to download these BUT from the UK archive, not the US archives thanks to alternate links provided, for example:
curl -L ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR319/005/SRR3191545/SRR3191545_1.fastq.gz -o SRR3191545_GSM2073124_ZIKV2-1_Homo_sapiens_RNA-Seq_1.fastq.gz
There is no explanation anywhere that I could find that explains the disappearance of folders from the FTP site…
P.S. I found the SRA-explorer thanks to an entry on this discussion link: https://www.researchgate.net/post/What_is_fastest_way_to_download_read_data_from_NCBI_SRA
specifically:
Appendix:
- Docker Hub entry maayanlab/zika
Pingback: Dockerization – details needed – Biochemistry Computational Research Facility (BCRF) – UW–Madison
It seems quite handy. Thanks, Jean-Yves!